What do we do till thenAnd NO vaccine at least for next couple of years.
Nobody said otherwise.What does he say ? NON ICU WILL MAKE A FULL RECOVERY
But what does he say about who will land up in ICU and who won't?
What do we do till thenAnd NO vaccine at least for next couple of years.
Nobody said otherwise.What does he say ? NON ICU WILL MAKE A FULL RECOVERY
What do we do till then
 theprint.in
theprint.in
Absolutely!! If RT PCR is not feasible, use serology to calculate the incidence and mortality rate.This is kinda the only real option:

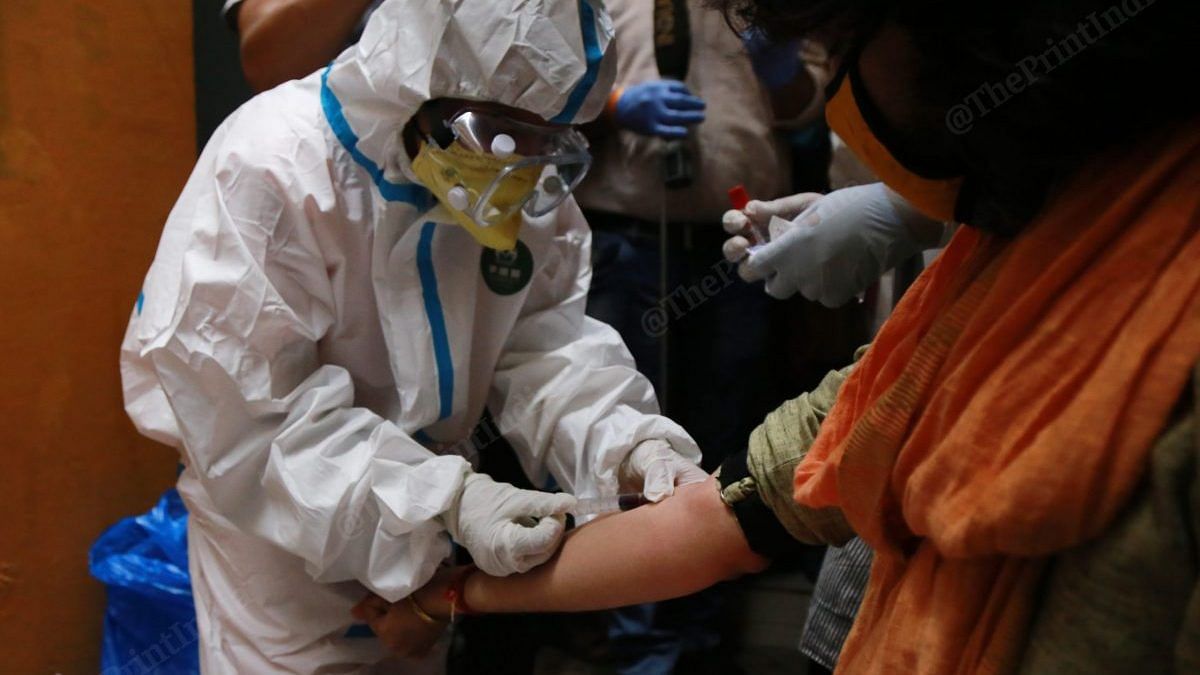
Lift lockdowns, protect the vulnerable, treat Covid like a health issue and not a disaster
Serological studies have shown that 10 times more Indians have had Covid than reported, and the death rate is 0.1%. So let’s not exaggerate, say experts.theprint.in
Watching Prime Time with Ravish Kumar. Today's episode is awesome. Lolz.
Finally saw it today after dilly dallying.. must say he roasts B n D media like a pro. It's his suttleness that makes him stand apart from other paid media.Watching Prime Time with Ravish Kumar. Today's episode is awesome. Lolz.
Yup, he comes from Hindi heartland, so proficiency is Hindi is needed to get his point. Even I found it hard at times to get hold of his sarcasm and serious reporting.Tried watching it, but anything more than street slang bambaiya hindi goes over my head. Couldn't understand 90% of it. He seems a bit like John Oliver though.
Since it is probably going to be my last post about Covid 19 in this forum, let me try to explain the very limited (and often flawed) role of R naught value.
The established definition of R0, as phrased by Anderson and May, is “the average number of secondary infections produced when one infected individual is introduced into a host population where everyone is susceptible”. They have stated that “If R0 is greater than one then the outbreak will lead to an epidemic, and if R0 is less than one then the outbreak will become extinct”
Sounds simple and logical but is it really? ( they have assumed that R0 is a threshold parameter that establishes whether an outbreak yields an epidemic or not)
Here I will try to establish that the average number of secondary infections (i.e., R0) is not always an epidemic threshold parameter without going deep into statistics.
Epidemiologists calculate R0 using individual-level contact tracing data obtained at the onset of the epidemic. Once an individual is diagnosed, his/her contacts are traced and tested. R0 is then computed by averaging over the number of secondary cases of many diagnosed individuals. This approach is based upon the definition of R0, but it does not ensure that the calculated R0 is also an epidemic threshold parameter.
Another approach (which is more commonly used) is to obtain R0 from population-level data, namely cumulative incidence data. Making certain individual-level modeling assumptions (e.g., the mass-action principle of infectious spread, time independent infection rates, etc.), theorists construct models (typically) based on Ordinary Differential Equations (ODEs) which describe the dynamics of the expected population size in different disease stages without tracking individuals.
It is very important to note that the individual-level modeling assumptions cannot be verified using population-level data (i.e., they remain hypothetical). ODE models are formulated in terms of disease transmissibility and progression rates at the population level. These parameters are obtained by fitting the model to population-level data; their relation to the individual-level processes may be quite complex and is generally unknown.
Bifurcation analysis of the ODE model yields a threshold parameter , that signals the epidemic as indicated by Anderson and May and is formulated in terms of the population-level parameters. This threshold parameter is not usually checked against the value of R0 that has been calculated from contact tracing data !!
Now let`s take a very simple example,
A simple ODE model is the Susceptible-Infected (SI) model given by dS/dt = π-βIS/(S+I) and dI/dt = βIS/(S+I)-μI, where β and μ are the inflow and, respectively, the outflow of infectious individuals per infectious capita. We apply this model at disease invasion when virtually everyone is susceptible (i.e., S/(S+I) is approximately 1) and obtain dI/dt = βI-μI. The threshold parameter for the reduced model is β/μ; if β/μ>1 an outbreak develops into an epidemic, if β/μ<1 an outbreak goes extinct.
It is important to note that β and μ are obtained from fitting the model to population-level data, with no clear association to the causal individual-level processes. An individual-level model that is compatible to these dynamics is a branching process ( I will not get into the branching process, as it will make things slightly complicated. However, the branching process is not the only possible ILM that is compatible with the ODE model. I can show that other plausible ILMs can be constructed very easily that yield the same ODE dynamics as the SI model at disease invasion.
In short, certain population-level dynamics, theoretically specified by an ODE model, can be the result of many distinct ILMs. I can easily demonstrate that the R0 obtained from the ILM, by applying the definition of Anderson and May may be different from the epidemic threshold parameter provided by the ODE model. Therefore, population-level predictions based upon an ODE model that use the R0 value found by contact tracing as a threshold parameter may be (in reality most of the time) will not only be inaccurate, but also can have disastrous outcomes if containment plans are modified based upon it.
Thus, measuring R0 through contact tracing (as generally occurs during an outbreak investigation), may not help in predicting the severity of the outbreak and may not be a useful measure for determining the strength of the necessary control interventions. Only an epidemic threshold parameter can be used to design control strategies. This parameter can be obtained through fitting an ODE model to population-level data.
Hope this post does not sound cynical, it is just basic epidemiology and statistics. R naught is great for movies on epidemics, but it is a very outdated concept and Anderson himself wrote an extensive article couple of years back that his theory is not to be taken in to account during containment program during an active epidemic.
